Welcome to LARKSdb
LARKSdb is meant to be a useful tool to predict regions of proteins that may be involved in reversible amyloid formation and phase-separation based on protein structure predictions. Threading and 3D profiling are used to predict if a 6-residue protein segment can form a LARKS structure.
How to read the protein graph
 A grey bar means the following six-residue segment is predicted to form a LARKS structure. Most segments are not favorable to form a LARKS structure which is why not every segment has a corresponding bar on the graph.
A grey bar means the following six-residue segment is predicted to form a LARKS structure. Most segments are not favorable to form a LARKS structure which is why not every segment has a corresponding bar on the graph.
The magnitude of the bar represents how confident the prediction is based on predicted energy of formation by a Rosetta algorithm, but for most purposes these charts are best interpreted in binary: either a segment is predicted to form a LARKS, or it is not.
The text at the top is the protein sequence. Lower-case letters correspond to low-complexity sequences as highlighted by the SEG algorithm.
What to look for in the protein graph
Phase-separation to liquid or solids requires multivalency. A protein must have multiple exposed LARKS to interact with to phase-separate. Therefore look for regions with several predicted LARKS in close proximity. LARKS must be exposed to interact, so look for LARKS in predicted disordered or low-complexity regions.
Comparison to other amyloid
Amyloid fibrils are usually associated with stability, but fibrils formed by LARKS rich proteins are often labile. Below two graphs are shown as representatives of solved fibrils from the disease-related protein tau (PDBID: 5O3T) and the labile amyloid FUS (PDBID: 5W3N). The profile for tau has few predicted LARKS segments where the FUS graph is replete with LARKS.
FUS fibril example
LARKS profile
|
Short regions of beta-sheet structure highlighted in orange. Beta-sheets are very short and often interrupted by kinks leading towards a less stable structure.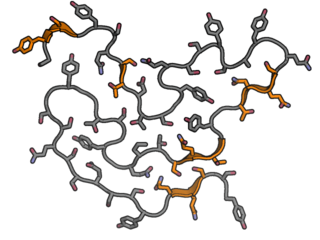 |
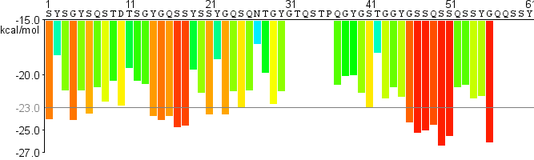 zipperDB profile |
TAU fibril example
Tau has long beta-sheets formed from heto-zippers. Tau fibrils are extremely stable and resistant to denautration, unlike FUS fibrils.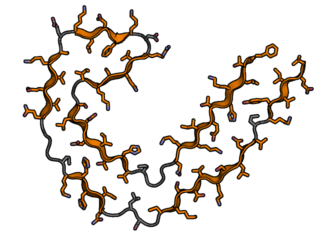 |
LARKS profile
|
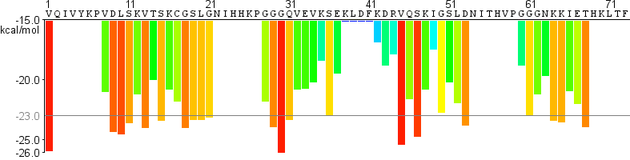 zipperDB profile |